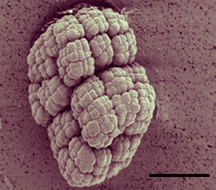
K. radiotolerans typically grows as symmetrical, multi-cell clusters (approximately 10µm), though individual round cells (1-1.5µm) expressing a single polar flagellum have been observed under certain conditions. Cell growth occurs between 11-41∞C, pH 5-9, and in the presence of < 5% NaCl and < 20% glucose. Physiological assays have revealed a seemingly straightforward metabolism consisting primarily of carbohydrates and alcohols as growth substrates. More than 90% of cellular fatty acids are anteiso 15:0, but arachidonic acid and 19-24 –C chain length alkenes (approximately 70% were 21- and 22-C chain length alkenes) are also produced. Kineococcus radiotolerans 16SrRNA gene sequence shows 93% identity to K. aurantiacus but whole genome hybridization revealed only 31% similarity, indicating significant genomic disparity. One characteristic feature of K. radiotolerans is the dramatic change observed in colony morphology with age on plating medium. Young colonies are moist, smooth and round, but transition into a rough, dry, raised colony mass of irregular shape over prolonged incubation. Mature colony morphology is strikingly similar to that of Mycobacterium tuberculosis. TEM of K. radiotolerans thin sections revealed a thick extracellular polymer shell surrounding individual cells within cluster formations. Related K. aurantiacus lays down an extracellular polymer consisting of meso-diaminopimelic acid and arabinogalactan. Kineococcus radiotolerans exhibits a high degree of resistance to ionizing g-radiation, as well as desiccation, keeping with the proposed model of a highly efficient cellular defense and repair system for Deinococcus radiodurans (Mattimore and Battista, 1996). Stable population sizes were observed following Co60 dosing experiments up to 3.5 kGy and less than 1 log difference in survivorship was measured compared to Deinococcus radiodurans. Not unlike Deinococcus radiodurans, K. radiotolerans retains viability following long-term desiccation. Kineococcus-like 16S rRNA gene sequences have been reported from the Mojave desert (Garrity and Searles, 1998) and air samples collected in the northern Caribbean following African dust storm events (Griffin et al., 2003), suggesting Kineococcus-like microorganisms may flourish in arid environments. Because of its remarkable resistance to ionizing radiation and desiccation, K. radiotolerans has potential usefulness for in situ biodegradation of problematic organic contaminants from highly radioactive environments. Moreover, completion of whole genome sequence from other radioresistant microorganisms Deinococcus radiodurans and Rubrobacter xylanophilus presents exciting opportunities for functional and comparative genomics explorations into strategies for survival in high radiation environments and determination of the evolutionary origins of radioresistance mechanisms and highly efficient DNA repair machinery. Relevant Publications Phillips, R.W., J. Wiegel, C.J. Berry, C. Fliermans, A.D. Peacock, D.C. White, and L.J. Shimkets. (2002) Kineococcus radiotolerans sp. nov., a radiation resistant, Gram-positive bacterium. Int. J. Syst. Evol. Microbiol. 52:933-938. Garrity, G.M., and D.B. Searles. (1998) Direct sequence submission to the NCBI GenBank database. Griffin, D.W., C.A. Kellogg, G.A. Garrison, J.T. Lisle, T. Borden, and E.A. Shinn. (2003) Direct sequence submission to the NCBI GenBank database. |
||
|
||
Kineococcus radiotolerans