Photo credit:
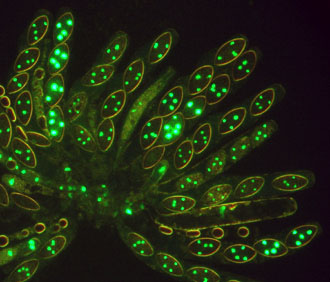
Namboori B. Raju, Stanford University. Neurospora tetrasperma . A rosette of maturing asci from
a cross of histone H1-GFP x wild type. Asci are mostly four-spored
and the ascospores are heterokaryotic for hH1-GFP. Histone H1 is
a chromosomal protein, which is synthesized in the cytoplasm and
incorporated into all nuclei in the shared cytoplasm. Thus all four
nuclei (two hH1-GFP and two non hH1-GFP) of each ascospore glow equally
bright. See, Jacobson, D. J., Raju, N. B., and Freitag, M. 2008.
Evidence for the absence of meiotic silencing by unpaired DNA in Neurospora
tetrasperma. Fungal Genetics and Biology 45:351-362.
|
Neurospora is
the premier model for filamentous fungi. Additional Neurospora sequenced
genomes, combined with the existing N. crassa and more distant
outgroup species sequences, will create an environment for genomic comparative
biology that will be unsurpassed for studies of genome evolution. N.
tetrasperma, although closely related to N. crassa, is
the only Neurospora species that employs a mating strategy
that combines selfing with occasional outbreeding using a novel genetic
system,. This system, known as pseudohomothallism is
novel to fungi. N. tetrasperma provides a model for
studying both the genomic and evolutionary consequences of asexuality
and inbreeding, two mating strategies that are widespread among plant
pathogenic and animal pathogenic fungi. These consequences cannot
be studied in obligate outbreeders alone, such as, N. crassa and
other popular genetic models.
Among
the many new research area that will be enabled by access to multiple Neurospora sequences
include adaptation of fungi that interact with forest trees; genome
defense against mobile elements; evolution and genetic consequences
of the widespread, but unique, fungal mating strategy pseudohomothallism. Forming
a strong foundation for new research in evolutionary biology and
ecology stands a large and strong genetic community with an NSF funded
Fungal Genetics Stock Center containing thousands of mutants and
more than 5000 wild type isolates from global collecting efforts. Neurospora also
offers numerous advantages for basic research in genetics, molecular,
cell, and evolutionary biology. The genetic model species Neurospora
crassa has a sequenced 43 mb genome that
contains 10,620 predicted genes - a complexity comparable to that of
animal model systems such as Drosophila. Not itself a pathogen,
it has long served as the model of choice for the estimated 1.5 million
filamentous fungal species, which include important animal and plant
pathogens. In terms of agroforestry, diseases of plants caused by filamentous
fungi are guaranteed to become increasingly important and, therefore,
will adversely affecting biorefining. On the positive side,
filamentous fungi also are essential to the decay of plant material
for the production of biofuels and bioproducts, particularly ethanol
and organic acids.
|